
4 minute read
There are billions to trillions different proteins in Nature (20,000 protein gene recipes in the human body) and each has a specific fold or shape. The shape determines its function. Proteins’ 3-D structures create specific binding sites that only fit certain targets, like a lock and key. Knowing their shapes is like conceptualizing the building blocks of Nature, of biological entities.
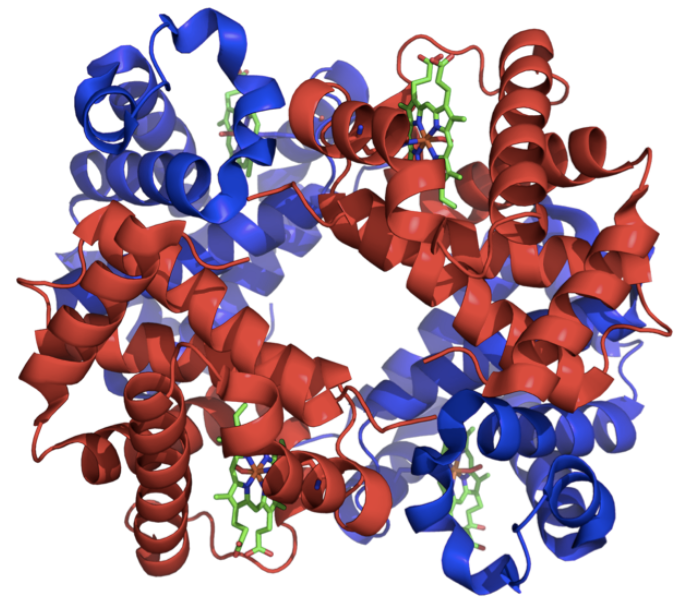

Proteins are made up of amino acids (between roughly 100-1000, 300-400 = average per protein). What determines the particular shape of a protein, how it folds, the Lego piece it becomes, are the chemical interactions between the amino acids. For example, two chemical reactions between amino acids include hydrogen bonds (positive-negative attraction, like a magnet) and hydrophobic effects (molecules avoiding water and clustering together).
The incredible complexity of amino acid chains’ chemical interactions has made the conceptualisation/capturing of proteins’ shapes extremely challenging. Before DeepMind/A.I./AlphaFold, scientists used a painstaking process called X-Ray crystallography to do this, allowing researchers to accurately pinpoint the shapes of only 100,000 protein structures out of the billions in Nature.
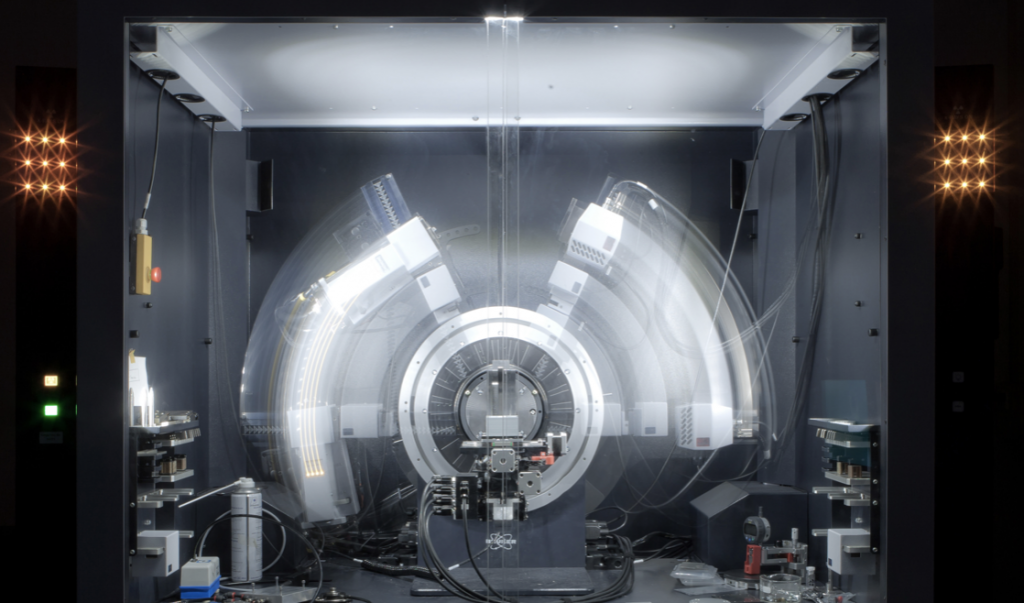

X-Ray crystallography, which takes weeks to months to years per protein shape and costs $50,000-$500,000, depending on the protein’s complexity, works like this:
1.) Grow a crystal (purified protein is placed in special conditions, molecules arranged in a repeating crystal lattice…the repetition is essential since it amplifies the X-Ray signal when it hits it).
2.) Shine the X-Rays on the crystal (electrons in the atoms scatter the X-rays, creates a diffraction pattern/spots on a detector). Note: there are only 16 synchrotron facilities (which provide much brighter, more tunable, more precise X-rays than typical machines) in the U.S. and they cost $100 million to $1 billion to build.
3.) Record the diffraction pattern (thousands of measurements are made from different angles).
4.) Mind-boggling math and compute (Fourier transforms, solve the phase problem to recover missing information, produce an electron density map/a 3-D cloud of where the electrons are).
5.) Build a protein model (fit the amino acid sequence into the density map, adjust and refine the model until it matches the data).
Scientists went through with this labor-intensive, capital intensive, mentally-taxing, complex process (sometimes thousands of crystallization trials, proteins are fragile, flexible, and hard to crystallize) because knowing protein shapes is super helpful and allows us to:
1.) Design drugs that fit exactly into target proteins
2.) Block harmful proteins or activate beneficial ones
3.) Understand diseases (misfolded proteins are responsible for Alzheimer’s, Parkinson’s, Sickle Cell Disease, etc.)
4.) Predict Biological Behavior (like will my left nipple keep burning when I drink grapefruit juice)
5.) Biotechnology and engineering (improve industrial processes/enzymes in food production or detergents)
If we could create a database of all proteins’ shapes, this would open many, many doors for medical, industrial, and biological breakthroughs:
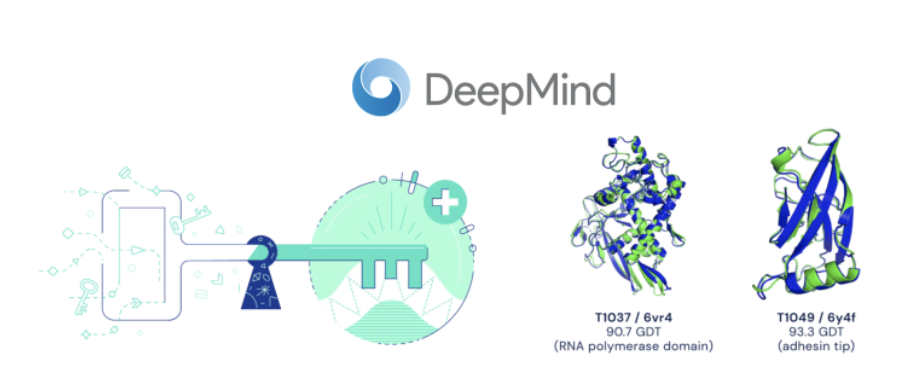
Enter David Baker, Demis Hassabis, and John Jumper:

Scientists like to reduce the complexity of Nature into simple, mathematical terms. But biology was perhaps too messy and emergent to be captured in terse mathematical statements (264, The Infinity Machine). Scientists also don’t like black box tech and prefer transparency in every step of the research method, but to Jumper, opaque models that gave you an answer were better than transparent ones that failed (264, The Infinity Machine).
264: The effort to understand biology through the axioms of physics was a dead end…
Enter Uniprot Dataset and A.I.:
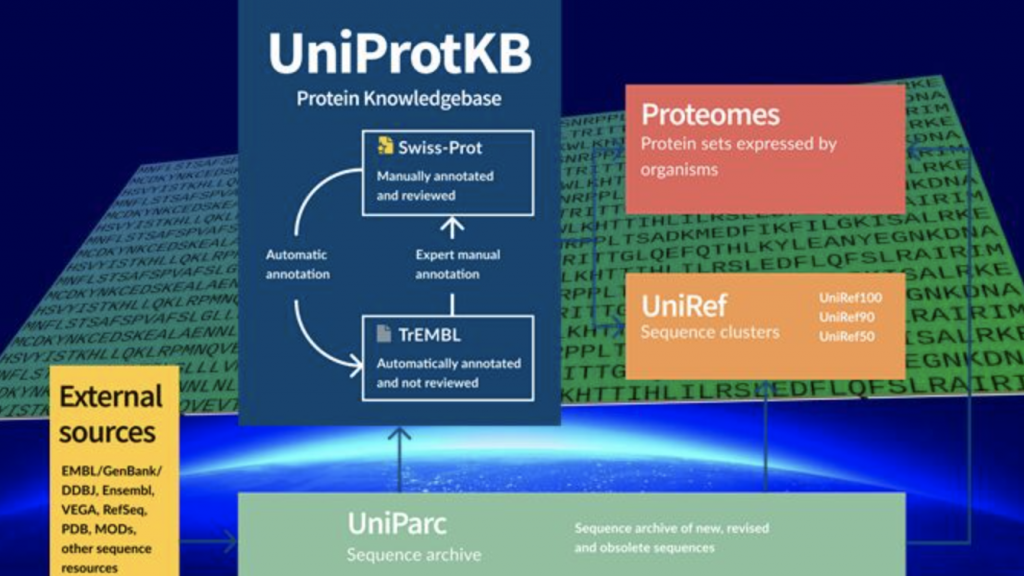
While protein shapes are a hassle to capture, discovering the sequences of amino acid chains is much, much easier. A simple chemical process (Edman degradation) had allowed researchers to not only document the 20,000 human protein amino acid sequences, but also millions of amino acid sequences of plants, animals, fungi, and bacteria.
In addition to analyzing the available protein shapes, AlphaFold would deeply analyze this Uniprot Dataset. Of course humans are not plants nor fungi, but we share the same, basic biological building blocks, so “understanding” fungi amino acid sequences could give the A.I. some opaque, predictive power. Basically, DeepMind borrowed ideas from structural biologists: amino acids that appeared in nearly all the chains in a particular kinship group were assumed to play an important role in the resulting protein structure. Some amino acids had evolved in pairs (267). This long, evolutionary history had hidden patterns and fundamental rules that A.I. (like it had done for the game, Go) could discover, which humans would likely never have grasped, even after centuries of analysis and crystal X-Rays.
But here is the incredible, goosebump-inducing breakthrough: The standard approach, for scientists interacting with the Uniprot Data set, was to create a contact map of predicting which acids in an amino acid sequence would touch each other in the folded protein structure (268). This is good, but AlphaFold could do better:
AlphaFold predicted the exact distances between each amino acid. The shift from a clunky and crude contact map to a precise distance map, or distogram, was like going from black and white to a full color TV (268).
Then, using a transformer (the ‘T’ in ChatGPT), the AlphaFold model ingested the entire UniProt database, teasing out the meaning in the evolutionary patterns (274). AlphaFold also trained itself, feeding the model’s own protein structure predictions back into its training set.
AlphaFold saw the patterns. The protein shapes were discovered. Pop the champagne.

DeepMind catalogued the shapes of all 20,000 proteins in the human proteome, 83% of which had not been mapped by crystallography (277).
In the summer 2021: AlphaFold plotted 350,000 protein structures, from yeast to fruit flies. By July 2022: AlphaFold had folded around 200 million proteins in total (277).
Applications include proteins digesting plastic in the oceans, crops resisting disease without pesticides, and accelerated drug discovery (277).
The protein mapping field was astounded: most scientists believed this feat was never going to be accomplished in their lifetimes. The average chain of amino acids could theoretically be twisted into 10^300 possible shapes – trillions upon trillions of forms. Yet AlphaFold discovered the correct shapes of folded chains, without the help of superpositions or quantum computing.
In 2024, John Jumper, Demis Hassabis, and David Baker won the Nobel Prize in Chemistry. Unfun fact, I asked ChatGPT: when did Demis Hassabis and John Jumper win the Nobel Prize, and it firmly denied their winning the prize:
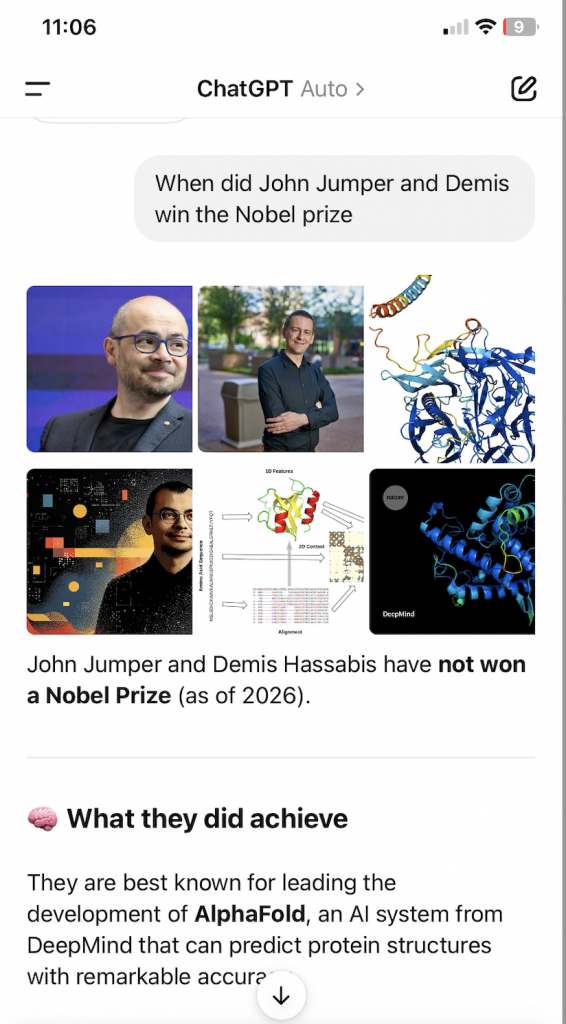
Competition, conspiracy, or hallucination?
So yes A.I. has its dark side, addicting us on social media, creating slop, shouldering out artists, gripping the mentally unstable via chatbots and prodding them towards heinous acts, providing us with a tempting, potentially-degrading, mental crutch, upheaval in work, occupational-existential-abysses, and persuasively expressing false knowledge, but we should also try and remember and champion the breakthroughs, shift and adapt (because let’s be honest: it’s here to stay), and harness the discoveries.
